
Analysis of Barry Hall’s Research of the E. coli ebg Operon
Understanding the Implications for Bacterial Adaptation to Adverse Environments
Abstract
Much research has been done on the ebg operon of the bacterium Escherichia coli over the last 30 years. specific mutations within this operon enable the bacterium to metabolize lactose.
Keywords: Mutation, Adaptive, ebg, Hall, lac, Hypermutable, Evolution, Natural selection, Escherichia coli, operon, Bacteria
This paper was originally published in the Proceedings of the Sixth International Conference on Creationism, pp. 149–163 (2008) and is reproduced here with the permission of the Creation Science Fellowship of Pittsburgh (www.csfpittsburgh.org).
Abstract
Much research has been done on the ebg operon of the bacterium Escherichia coli over the last 30 years. Although the function of the ebg operon is still unknown, it has been observed that specific mutations within this operon enable the bacterium to metabolize lactose sufficiently to allow growth. Interestingly, this growth occurs in a lacZ- genotype (gene for ß-galactosidase in the lac operon). Thus, this gene has been referred to as an “evolved ß-galactosidase”, and has been widely accepted as an example of “evolution in action”. Under these cultivation conditions, the ebg operon appears to harbor adaptive mutations. Mutations (at codons 92 and 977) in the ebgA gene (which codes for ebg ß-galactosidase) occur consistently when an E. coli lacZ- population undergoes carbon starvation in the presence of lactose. These are the only mutations observed in the ebgA gene and these particular mutations are not found when the bacteria are subjected to different adverse environmental conditions. Mutations are also found in other genes suggesting a mechanism which has affects on the entire genome. Several models have been proposed to explain this phenomenon.
Hall’s work needs critical evaluation. Mutations in the Ebg system are clearly not an example of evolution but mutation and natural selection allowing for adaptation to the environment. Several possibilities for the function of the Ebg system are suggested. In addition, there is an assessment of the likelihood of these mutations in the ebg operon occurring in a natural setting. An implication of this research is an understanding that adaptive mutation makes “limited” changes which severely restrict its use as a mechanism for evolution.
Adaptive mutations can readily fit within a creation model where adaptive mechanisms are a designed feature of bacteria. Further understanding of these mutations in the ebg operon may help the development of a creation model for adaptation of bacterial populations in response to the adverse environmental conditions in a post-Fall, post-Flood world.
Introduction
For over 150 years the mantra of evolutionists has been that random changes/mutations directed by natural selection is the driving force of evolution (common descent of all life). Although biblical creationists do not deny that these forces occur, they do not equate them with evolutionary transformations necessary for common descent. Edward Blyth, a creationist, was one of the first individuals to formulate the idea of natural selection. Mutation and natural selection in populations of bacteria, animals, and humans are readily observable. Mutations can be classified as deleterious, neutral or beneficial with beneficial being by far the smallest category. Beneficial mutations typically lead to relatively small changes that give an organism an advantage in a specific environment. For example, in a population of beach mice (Peromyscus polionotus) a mutation was found in the Mc1r gene that alters the coat color such that it is lighter, similar to the color of the sand where the mice live.1 In the beach environment, there is a selective advantage for this lighter coat color because it may allow better camouflage protection from predators. However, the benefit of this mutation is extremely limited in its scope. Should the environment change or the mouse population relocate, this particular allele of the Mc1r gene might become disadvantageous, making the value of this beneficial mutation very context-dependent. Also, the light fur coloring resulted from a mutation that decreased pigment production. Hence, it may have provided a beneficial phenotype, but the mutation was genetically deleterious.
blockquote class="pull right">Another example of beneficial mutations is antibiotic resistance in bacteria.Another example of beneficial mutations is antibiotic resistance in bacteria. This has become a popular example of “evolution in action”. Antibiotic resistance mutations encompass a wide variety of mutations that alter targeted bacterial proteins. For a comprehensive review of these see.2 Although the mutant bacteria gain a survival advantage in the presence of antibiotics, they often do not compete as well with wild-type bacteria in an environment without antibiotics. In addition, resistance may provide a beneficial phenotype, but it usually results from mutations that eliminate or reduce specific genetic function, such as gene regulation or expression.2 So again, this type of change is beneficial only in very specific environments.
Mutations are commonly thought to be random changes in the DNA that may or may not be beneficial to an organism in its environment. However, studies have indicated that adverse environmental conditions may initiate mechanisms in bacteria that lead to mutations which specifically allow the bacteria to survive and grow in the given environment.3,4,5,6 Initially these were termed “directed” mutations but are now more appropriately called adaptive mutations. Barry Hall, professor emeritus of the University of Rochester, New York, has done extensive work in the field of adaptive mutations. His work on the ebg operon in Escherichia coli has been offered to refute biochemist Michael Behe’s7 challenge that some systems have a complex interrelationship with no possible evolutionary intermediate stages or transitions of development (that is, irreducible complexity). In his book, Finding Darwin’s God, Kenneth Miller believes,
The most direct way to do this [show irreducibly complex systems can evolve] would be with a true acid test—by using the tools of molecular genetics to wipe out an existing multipart system and then see if evolution can come to the rescue with a system to replace it.8
Miller is convinced that Hall’s work with the ebg operon passes the “true acid test” with flying colors, “No doubt about it—the evolution of biochemical systems, even complex multipart ones, is explicable in terms of evolution.”8 Behe and Miller have had several internet exchanges devoted to this topic.9, 10 While mutations can help bacteria adapt to their environment, such mutations should not be equated with evolution.
There are two reasons why the concept of adaptive mutations would appear to be problematic for neo- Darwinism proponents. First, although the mutations themselves are random (at least according to current models), the mechanism for generating mutations is specifically responding to environmental clues. In this setting, mutation and natural selection become goal-oriented. While this is expected for an adaptation mechanism, evolutionary mechanisms are specifically identified as goal-less. A second reason is that the mechanisms of adaptive mutations may reveal there are limits to the genetic change possible by mutations of genes in organisms. With the ebgA gene, only two mutations in the gene were ever observed after countless in vivo studies. On-the-other-hand, in vitro studies have shown a wide variety of mutations that can make the enzyme more efficient in hydrolyzing a substrate.11
It may be especially important for bacteria to genetically possess a “rigid flexibility.” Such flexibility would allow bacteria the necessary genetic versatility to adapt to exposure to ever-changing environments.12 Yet, the rigidity of possible changes would limit the bacteria to remain within its created baramin.13 This fits with observable evidence of bacterial changes. The study of adaptive mutations is important for understanding the power and the limitation of mutations for change in bacterial species. This may assist creation models of speciation, adaptation, and pathogenesis of bacteria in the post-Fall and post-Flood world.
Definition of Adaptive Mutation
Hall summarizes adaptive mutations well when he writes:
Adaptive mutations differ from growth-dependent mutations in two key respects. First, adaptive mutations occur in nondividing or slowly dividing cells which are under selection for a particular phenotype, whereas growth-dependent mutations occur in dividing cells that are not under strong selection. Second, adaptive mutations produce only those phenotypes which allow the cells to grow, whereas growth-dependent mutations occur randomly with respect to their effects on fitness.14
Adaptive mutations are those that allow bacteria to survive preferentially under non-lethal selection. These mutations were once called “directed” as they were thought to occur only in genes that allowed for the metabolism of the metabolite being used in the selection. However, adaptive is a better description as mutations in genes not under selection have also been found.6 The bacteria used for studies of adaptive mutation are typically deficient for the ability to metabolize a particular substance, like lactose.
Initially, metabolically deficient bacteria (auxotrophs) are grown on minimal media supplemented with the metabolite (that the bacteria cannot metabolize) as the sole energy source. Most of the bacteria do not grow on this media. Those that grow shortly after selection begins (usually within two days) most likely have mutations that allowed them to utilize the metabolite before they were plated on the minimal media. These are growth dependent mutations that occurred during the normal process of growth of the bacteria on complete media before placement on minimal media. However, 95% of the mutations that occur once the bacteria have been cultivated for a week on minimal media are adaptive mutations that occurred after being placed on the minimal media.15 This situation of starvation closely mimics reality for those bacteria living in nutrient poor environments.
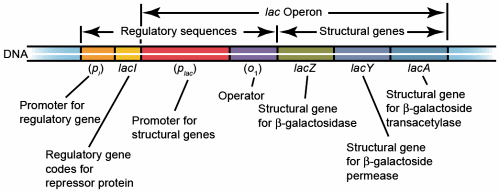
Fig. 1. Lac operon. Additional operator region O2 is located upstream of O1 and operator region O3 is located within lacZ.
In some cases adaptive mutations in bacteria are simple reversions of the auxotroph back to the prototroph. For example, one Lac- strain of E. coli contains a + 1 frameshift in the lac operon (on the F' plasmid) which inactivates it. Adaptive mutations in this strain are a reversion of the frameshift such that the lac operon is now active.16 Hall’s research involved using an E. coli Lac- strain (W4680) in which there is a deletion of approximately 1/3 of the lacZ gene.17, 18 This cannot be corrected through a simple reversion in the lac operon, thus adaptive mutations were not seen in the lac operon but in the related ebg operon.
Lac and ebg Operons
The lac operon of E. coli is subject to negative regulation. It consists of several genes designed to allow lactose into the cell, for the catabolism of lactose, and the regulation of the lac operon (fig. 1). lacZ, Y, and A are structural genes encoding ß-galactosidase (ß-gal), galactoside permease (lac permease), and thiogalactoside transacetylase, respectively. lac ß-gal is responsible for the catabolism of lactose and the isomerization of lactose to allolactose needed to induce the lac operon. Lac permease is a membrane bound protein that transports lactose into the cell. Thiogalactoside transacetylase is encoded by the lac operon, but its function is not known. lacI encodes the lac repressor. A promoter and operator sequence are located between lacI and genes lacZ, Y, and A. When lactose is absent, the repressor binds to the operator and greatly reduces the rate of transcription of lacZ, Y, and A. This is an energy conservation mechanism since E. coli preferentially catabolizes glucose. In the absence of glucose and presence of lactose, lactose is isomerized to allolactose by lac ß-gal. Allolactose binds to the lac repressor which prevents it from binding to the operator and transcription of the lac operon structural genes greatly increases. Isopropylthiogalactoside (IPTG) is a powerful, nonmetabolizable inducer of the lac operon commonly used in experimental studies.
The ebg operon of E. coli (fig. 2) is also subject to negative regulation19 but the natural substrate of this operon is unknown.20 ebgA, B, and C are structural genes encoding the a subunit of the ebg ß-galactosidase (ebg ß-gal), a protein of unknown function but structurally similar to transport proteins, and the ß subunit of ebg ß-galactosidase, respectively.20 ebgR encodes the ebg repressor.20 The promoter and operator sequences have not been determined experimentally but have been predicted from the DNA sequence.21 ebg ß-gal can catabolize lactose (better than any other ß-galactosidase sugar) but not sufficiently to allow growth on lactose even when approximately 5% of the soluble protein in the cell is ebg ß-gal (due to a defective repressor).20 Mutations in both ebgA and ebgR are required for lactose utilization.22

Fig. 2. Proposed structure of ebg operon.
lacZ and ebgA sequences show a 50% DNA homology and a 33% amino acid homology.23 lacI and ebgR sequences show a 44% DNA homology and 25% amino acid homology.24 Stokes and Hall have hypothesized that rather than gene duplication these two operons may have been the result of whole genome duplication sometime in the distant past. Supposed evidence for whole genome duplication has been shown for several species including E. coli and Sacchromyces cerevisiae.25, 26 However, this is a highly speculative reconstruction and totally based on evolutionary relationships. In addition, one might expect the mutational spectrum to be similar if the lac operon is the ancestor of the ebg operon or vise versa. The mutational spectrum of lacI and ebgR has been shown to be completely different with transitions (that is, AT replacing GC) dominating lacI mutations and transversions (that is, TA replacing GC) dominating ebgR mutations.27 Further arguments against gene duplication as a mechanism for evolution that would also apply to whole genome duplications have been discussed elsewhere.28
Classification of ebgA Mutations
Hall discovered several different classes of ebgA mutants using different selection schemes.18 All of the selections were performed using E. coli K12 strains (W4680) that have the lacZ- deletion and are ebgR- (constitutive).18 The media was supplemented with IPTG, which is necessary to induce the synthesis of lac permease. This permease is necessary for translocation of lactose into the cell.18 Five classes of mutants (Classes I, II, III, IV, and V) were obtained.18 Further analyses of these mutants have shown Classes I, II, and IV to be relevant for future studies of ebgA29 (table 1). Following selection with lactose Class I and II mutants were obtained. Class I had a growth rate of 0.45 on lactose and 0 on other sugars (all growth rates are in units of hours-1)18 (table 2). In contrast, Class II had a growth rate of 0.19 on lactose and 0.26 on lactulose.18 Class IV mutants were obtained by selection of Class I mutants with lactulose18 or Class II mutants with galactosyl-D-arabinose.30 Class IV mutants had a growth rate of 0.37 on lactose, 0.18 on lactulose and 0.13 on galactosyl-D-arabinose.18 Recombination studies showed that Class IV mutants were double mutants consisting of a combination of Class I and Class II mutations.30
Later sequence analysis of Class I mutants showed a G→A mutation at 1566 resulting in the substitution Asp-92→Asn.21 Class II showed a G→T or C mutation at 4223 resulting in the substitution Trp-977→Cys.21 As predicted by an earlier study, Class IV was found to have both Class I and Class II mutations.21 The majority of strains did not carry mutations in ebgC.21 Interestingly of all the strains that were analyzed, the overwhelming majority only had mutations in ebgA at 92 and 977 which are active site residues.21 Active site residues in ebgA were found to be homologous with ß-galactosidases from three proposed phylogenetic groups of bacteria (table 3). The Class I mutation changes the ebgA active site to consensus for that residue across all three groups.20 The Class II mutation changes the ebgA active site to consensus for that residue within its own phylogenetic group.20 Interestingly, the Class II mutation and the residue at 579 in ebgA differ from E. coli lacZ (but are consensus for ebgA’s proposed phylogenetic group).20
Table 1. Classification of ebgA mutations. Table 1 provides a summary of the wild-type ebgA, and classes of ebgA mutants including their mutants, their growth on various mediums, and their transgalactosylation activity (ability to convert lactose to allolactose and induce the production of lac permease).
| Enzyme | Mutation(s) | Lactose | Lactulose | Galactosyl-D-arabinose | Transgalactosylation Activity |
|---|---|---|---|---|---|
| Wild-type | — | — | — | — | — |
| Class I | Asp-92→Asn | + | — | — | — |
| Class II | Trp-977→Cys | + | + | — | — |
| Class IV |
Asp-92→Asn Trp-977→Cys |
+ | + | + | + |
Table 2. Growth rates of wild-type ebgA and ebgA mutant bacteria. Table 2 provides a summary of the growth rates of wild-type ebgA and ebgA mutant bacteria on various mediums.
| Enzyme | Lactose | Lactulose | Galactosyl-D-arabinose |
|---|---|---|---|
| Wild-type ebgA | 0 | 0 | 0 |
| Class I | 0.45 | 0 | 0 |
| Class II | 0.19 | 0.26 | 0 |
| Class IV | 0.37 | 0.18 | 0.13 |
|
* Growth rate reported in units of hours-1 * Data from Hall.18 |
|||
Table 3. Alignment of the active site amino acids of representative members of each of the three clades derived from phylogenetic analyses of bacterial ß-galactosidases.54 Highlighted are those of E. coli ebgA and E. coli lacZ.
| E. coli ebgA (A) | DHRHEHEMYEHWFDW | |
| K. lactis (A) | NHRHEHEMYEHWFDC | |
| Arthobacter sp. lacZ (A) | NHRHEHEMYEHWFDC | |
| T. maritima lacZ (B) | NHRHEHEMYEHWFNW | |
| S. salivarius lacZ (B) | NHRHEHEMYEHWFNW | |
| E. coli lacZ (C) | NHRHEHEMYEHWFNW | |
| E. cloacae lacZ (C) | NHRHEHEMYEHWFNW | |
| 1 | 2 | |
* Data from Hall.20
* 1 represents position 92 and 2 represents position 977 in ebgA
* A, B, and C represent clades from phylogenetic analyses of ß-galactosidases from Hall.20
Class IV was discovered to have an activity not found in Class I and Class II. Class IV has transgalactosylation activity that allows it to convert lactose to allolactose.31 Thus, Class IV does not require IPTG as an inducer of lac permease, and can now regulate the lac operon itself.31 It is the only class of ebg mutants that could survive in vivo.
Classification of ebgR Mutations
Hall discovered that ebgR mutations are more wide ranging than those of ebgA (for a complete list of mutations see Hall).27 All of the selections were performed using E. coli K12 strains (W4680) possessing the lacZ- deletion and were also mutant for ebgA (ebgA51+, Class II mutant).17 The media was supplemented with IPTG which is necessary to induce the synthesis of lac permease necessary for translocation of lactose into the cell.17 The ebgA51+ operon is under the control of wild-type ebgR and is inducible by lactose and lactulose but not sufficiently to allow survival on either sugar.22 Selection using lactulose recovered strains that had mutations in ebgR (designated with an “L” for lactulose induction) that allowed growth via induction by lactulose as well as several other ß-galactosidases.17 This seems to indicate an altering of the specificity of ebgR.21 The fold induction increase by these sugars was 10 to 90 fold greater than their induction of the wild-type ebgR.17 Mutations in the predicted sugar binding region were revealed through sequence analysis of several of the ebgR+L mutants.21 Interestingly, all of the ebgR+L mutations were inducible and not constitutive.17 This suggests a strong selection for non-constitutive mutations in ebgR in addition to ebgA.
Later studies revealed that the majority of mutations in ebgR are mediated by insertion elements IS1 and IS30.27 Interestingly, only about 4% of lacI mutations are mediated by insertion elements.27
Natural Function of the ebg Operon
Although the natural substrate of ebg ß-gal is unknown, it is important to speculate on its natural role. A search of the NCBI protein database shows that several species of Shigella (a close relative of E. coli) possess the genes for ebgR and the a subunit of ebgA. Shigella poorly utilizes lactose due to the absence of the lac operon.32 It is unknown for both E. coli and Shigella if the ebg operon is essential. It is also unknown if Shigella would acquire similar mutations in the ebg operon if it were placed under similar conditions that produced the adaptive mutations in the E. coli ebg operon.
It is impossible to know if the adaptive mutations at 92 and/or 977 in ebgA affect its normal function. The function of the ebg operon may not be essential or may be replaced by a functionally redundant system since these mutants grow well under laboratory conditions. However, since these mutations occur in the active site of ebgA it is likely to affect its function by either causing ebgA to lose specificity or change specificity. This is commonly observed with mutations in enzymes.33
Pseudogene
One possibility is that ebgA is a pseudogene. Pseudogenes were once thought to be rare in bacteria, however, recent work indicates they are common; encompassing 1–5% of individual bacterial genomes.34 Just as in eukaryotes, prokaryotic pseduogenes may arise from duplications of native genes or mutations in native single copy genes.34 Unique to prokaryotes is the possibility that pseudogenes arise from failed horizontal gene transfer events.34 Pseudogenes were once thought to be defunct relatives of known genes. While this may be the case for some, recent studies have shown some pseudogenes can be functional in gene regulation.35, 36 Hall believes that the high level of sequence conservation between ebg ß-gal and other lac ß-gals diminishes the possibility that ebgA is a pseudogene.20 As papers regarding the functionality of pseudogenes have only recently been published (last five years), it is possible that ebgA is a pseduogene with an unknown regulatory function.
Backup system
Another possibility is that the ebg operon represents a functionally redundant backup system for the lac operon. In many species from bacteria to humans, functional redundancy exists between certain genes. Functional redundancy has been a deterrent in minimal genome determination for microorganisms as a gene may not be classified as essential due to the presence of two genes that can perform the same function—delete one and the other still performs the necessary function.37 Additionally, even if a microorganism can survive with only one copy of genes for a particular function, it may not be as robust to mutations since it is now lacking a backup copy. Interestingly it has been shown that these backup systems are regulated in such a way that if the essential gene becomes mutated or inactivated the backup gene (responsive gene) becomes activated.38 For example, in the yeast Saccharomyces cerevisiae, Fks1 and Fks2 are involved in cell wall formation and are functionally redundant.39 When Fks1 becomes inactivated by mutation, Fks2 becomes up-regulated.39 Kafri, Levy, and Pilpel state,
We thus challenge the view that such redundancies are simply leftovers of ancient duplications and suggest they are an additional component to the sophisticated machinery of cellular regulation. In this respect, we suggest that compensation for gene loss is merely a side effect of sophisticated design principles using functional redundancy.38
Although wild-type ebg ß-gal is not functionally redundant to lac ß-gal, it is possible that the regulation of backup genes may involve activating mutation mechanisms (such as those leading to the hypermutable state in bacteria—see appendix). This would result in mutations in backup genes that lead to proteins that can perform the same function as the proteins from essential genes. Possibly ebgA serves as a backup gene for several essential genes and the most flexibility within the system is achieved by altering ebgA in accordance with the particular essential gene that has been inactivated.
Further support of the idea that the ebg operon is a backup system is implicated by the findings that adaptive mutagenesis of ebgR is regulated. Hall found that mutations in phoPQ genes decrease the number of adaptive mutations in ebgR by a factor of 6.14 He also found mutations in several other genes that increase or decrease the adaptive mutation rate of ebgR but these were not further investigated.14 Interestingly, the disruptions in phoPQ did not affect the adaptive mutation rate of other operons such as trp and bgl, thus, phoPQ appears to be specific for affecting adaptive mutagenesis of ebgR.14 In addition, disruptions in phoPQ did not affect the rate of growth-dependent mutations in ebgR, only the adaptive mutation rate.14 PhoPQ responds to adverse environmental conditions (such as nutrient deprivation) by directly or indirectly up-regulating the expression of many genes.40 This lends further support to the idea that when lacZ is mutated and lactose is the only carbon source, mechanisms (such as phoPQ) are initiated. This leads to mutations in the putative Ebg back up system that result in a functional ebg ß-gal that can utilize lactose and allow survival and growth of the bacteria.
Mutations in the Ebg System are not an Example of Evolution
Evolution (primarily neo-Darwinism) is currently the dominant paradigm in origin science. Within this context, scientists have diligently worked to assign an evolutionary role for adaptive mutations. This is exemplified by the title from one of Hall’s papers, “Experimental evolution of Ebg enzyme provides clues about the evolution of catalysis and to evolutionary potential.”20
However, the most widely accepted definition of evolution is simply “change of gene frequency over time.” This vanilla definition neither describes the types of changes nor the extent of these changes. As such, it offers little insight into either the mechanism or the function of evolution. It also does little to distinguish itself from a creation model of origins.
A more appropriate definition (and explanation) of evolution is “descent from a common ancestor” or “common descent”. This infers that all contemporary life is descended from a common ancestor(s) billions of years ago through the progressive modification of their DNA. However, papers dealing with the Ebg system ultimately provided little evidence in support of “common descent.” Rather, the observations of these studies were simply the adaptive aspects of bacteria, as the term adaptive mutations implies. Bacteria, facing adverse environmental conditions, activate mechanisms to increase the rate of mutation in the attempt to generate a variety of mutations, one class of which may give the population the ability to survive and grow under a given adverse condition (see Anderson12 for a more complete description of this characteristic).
Not surprisingly, the field of adaptive mutagenesis was initially met with very strong resistance. Early findings by Hall and Foster/Cairns seemed to indicate that the only mutations that were occurring were those specific for dealing with the selective pressure that the bacteria were under.41, 42 This would seem to contradict the randomness of mutations (or a mutation rate independent of the environment) required by Neo-Darwinian philosophy.43 Torkelson et al. comments,
Adaptive mutation provoked controversy about whether mutagenesis mechanisms exist that direct mutation preferentially to a selected gene, in a Lamarckian manner. The unorthodoxy of this suggestion led many to argue that adaptive mutations must not exist.6
As mentioned, adaptive mutations were originally called directed mutations because they seemed to be specific in relation to the selective pressure. Stahl dubbed them, “a unicorn in the garden”.44 Lenski, Slatkin, and Ayala state,
If the hypothesis of directed mutation is, indeed correct, it has onerous implications for bacterial genetics and, in particular, for the use of bacterial populations as model systems for the study of evolutionary processes.45
Several recent papers proposing alternate “random” processes have been countered by additional research supporting a more “directed” attribute of the adaptive mutation (see Anderson and Purdom).46 If, indeed, adaptive mutations are “directed” by environmental conditions, a glaring problem would be posed to any phylogenetic reconstruction that is dependent on DNA sequence differences. Hall comments,
If the probabilities of particular mutations are subject to environmental modulation, then the number of observed differences between two sequences may be completely unrelated to the time since they diverged.47
Thus, environmental pressures are speeding up changes in the DNA, the mutation rate has not been constant throughout time, and DNA sequence differences are not an accurate “clock”.
“A true acid test”
As mentioned previously, Kenneth Miller has used Hall’s work on the ebg operon as evidence that evolution can give rise to irreducibly complex systems.8 Miller believes that what Hall has observed is the “evolution” of a 3-part system necessary for lactose utilization. The three parts include a lactose-sensitive ebg repressor, ebg ß-galactosidase, and the ability of ebg ß-gal to form allolactose which induces lac permease. He states, “Unless all three are in place, the system does not function, which is, of course, the key element of an irreducibly complex system”.10
Once again, it is a matter of definition. How is Dr. Miller defining evolution? Adaptive mutagenesis of the Ebg system is not creating a new system, nor does it even impart E. coli with a new phenotype.Instead, the mutations, combined with selection, work on pre-existing genetic material. The wild-type ebg ß-gal already possessed the ability to catabolize lactose.20 The mutations merely restored the Lac+ phenotype that the mutant strain of E. coli had previously lost by a deletion of lacZ. In addition, mutations at 92 and/or 977 in ebgA simply allow the ebg ß-gal system to utilize lactose better. Although Class IV ebgA mutants (mutations at 92 and 977) can isomerize lactose to allolactose to induce lac permease, the Ebg system did not “evolve” the ability to transport lactose into the cell. The system is dependent on the presence of the lac permease from the lac operon. Thus, the Ebg system for lactose utilization is incomplete. Hall’s work with the Ebg system is not an example of the evolution of a new system or even a new phenotype. Rather the process of adaptive mutagenesis (through mutation and natural selection) has allowed the bacteria to make a minimal number of changes in pre-existing systems to regain a previously lost function in order to adapt to adverse environmental conditions. More time and more beneficial mutations as achieved by adaptive mutagenesis still do not result in evolution, merely adaptation.
Another problem for adaptive mutagenesis as a mechanism for evolution is the net overall loss of functional systems. As discussed in the appendix, the proposed hypermutable state of bacteria under non-lethal selection leads to genome-wide mutations.6 For example, in E.coli Lac- cells placed under non-lethal selection with lactose generated Lac+ mutants that also were Xyl- (inability to ferment xylose) or Mal- (inability to ferment maltose).6 Although these cells possessed beneficial mutations which allowed them to utilize lactose, they also possessed deleterious mutations that resulted in the loss of the ability to utilize xylose or maltose. Hall also found auxotrophic mutations among Trp+ revertants.42 It is not known if the mutations at 92 and 977 in ebgA affect the natural function of the Ebg system since its natural function is unknown. It is likely that these mutations do affect the natural function of ebg ß-gal since they occur in the proposed active site. Mutations in enzymes typically alter their substrate specificity or interfere with their ability to interact with their natural substrate.33
Hall, although not skeptical of evolution in general, does seem to conclude that his work on the Ebg system is not reflective of the experimental test or laboratory verification of evolution that Miller suggests. In a recent paper he recounted the history of his research on the Ebg system. Hall writes,
As a fresh young postdoc in 1972, I was pretty disdainful of evolutionary biology, dismissing it as just-so stories backed up by internally consistent, but experimentally untestable hypotheses.48
Underlying all of those questions was the big question: what did we have to know in order to predict both the evolutionary potential of an organism for a new gene function and the evolutionary potential of any particular ancestral gene?48
Well, we pretty much lost sight of that question over the years of studying Ebg. We had started out wanting to be able to predict evolution but ended up, much like classical evolutionists, simply explaining what we had seen. Sure we had watched the events occur in the laboratory, but no effort to predict outcomes in advance had been attempted.48
Of all living organisms, bacteria should be the perfect model for experimentally testing and verifying evolutionary predictions. Their quick generation time and large population size make it possible for laboratory observation of what have occurred over “millions of years” in nature.49 If mutation and natural selection can result in the net gain of functional systems, then this should be observable using bacteria in the lab. However, as Hall indicated, all he was able to offer was an explanation consistent with what he observed. He was not able to make predictions about how the organism would evolve. Bacteria (a subpopulation) when placed under adverse environmental conditions enter a hypermutable state that generates multiple mutations (see appendix). Some of these mutations are beneficial and some are deleterious. Selection favors bacteria that contain mutations that restore previously lost functions that will enhance survival under the current selective pressure. The same bacteria also have mutations that are not specific to the selective pressure and have resulted in the loss of other functions. Rather than being evidence for evolution which requires a net gain of new systems with new functions, this is clear evidence for an amazing phenomenon using pre-existing systems and pre-existing functions that allows for adaptation (with no overall net gain).
How applicable are Hall’s experiments to natural settings?
Although the mutations in the ebg operon have been observed to occur in a laboratory setting, could they also occur in a natural setting? One of our major challenges, also a concern of Behe9 concerning Hall’s work is the amount of “intelligent intervention” that had to occur to obtain a functional, regulated ebg operon that could utilize lactose. The following provides a summary of these challenges.
Use of IPTG
IPTG (isopropyl-beta-D-thiogalactopyranoside) is a synthetic analog of allolactose and induces the lac operon. IPTG was always present in his experiments (except where noted otherwise) to induce the production of lac permease needed to transport lactose into the cell.18 Since the ebg operon does not contain a gene for a lactose transporter, induction of the lac operon is still necessary. If IPTG is absent, lactose is not transported into the cell and subsequent mutations in ebgA and ebgR would not be positively selected.
Sugars used are not natural
Some of the sugars used for the selection schemes to obtain the different classes of ebg mutations are also not natural. Lactulose and galactosyl-ß-D-arabinose are not found in nature and lactobionate is rare in nature.48 The Class IV mutants (that can utilize lactose and form allolactose to regulate the lac operon) are only found after selection with either lactulose18 or galactosyl-D-arabinose.30 Therefore, it is hard to conceive how the ebgA mutations would have occurred naturally.
Double ebgA mutants cannot be obtained in a single step
Class IV mutants containing both Class I (at 92) and Class II (at 977) mutations were never obtained in one step.18 Class IV can be obtained either by selection of Class I with lactulose18 or selection of Class II with galactosyl-ß-D-arabinose.30 This is relevant as Class IV mutants are the only ones able to isomerize allolactose from lactose.31 A Class IV mutant obtained in one step would be necessary in a natural setting as IPTG would not be present and allolactose is necessary to induce the expression of lac permease needed for lactose uptake. Single mutants such as Class I and Class II alone would not be positively selected in a natural setting as these cells are not be able to induce the expression of lac permease. So the single mutants would not be available for further mutation and selection to give rise to double mutants. It appears that the only way the double mutant (Class IV) can be obtained is through an extremely artificial selection scheme involving the presence of a synthetic inducer and unnatural sugars.
Triple ebg mutants cannot be obtained in a single step
A completely functional, regulated ebg operon that can utilize lactose requires three mutations: one in ebgR and two in ebgA. The ebgR mutation could be constitutive, but preferably it would be one that allows regulation by lactose so that ebg ß-gal is only produced in the presence of lactose. This allows better economy and efficiency by the cell. The two mutations in ebgA (Class IV mutant) allow the cell to utilize lactose and form allolactose, which induces the expression of lac permease. However, all the selection schemes for ebgR or ebgA mutations always began with cells that were already mutant for one or the other. The ebgA mutations were obtained using cells that already had a constitutive ebgR mutation.18 The ebgR mutations were obtained using cells that already had mutations in ebgA that allowed it to utilize lactose and lactulose.17 Thus, no individual mutations in either ebgA or ebgR would be selected for. Mutations in ebgA without mutations in ebgR would not be selected for because ebg ß-gal would not be produced. Mutations in ebgR without mutations in ebgA would not be selected for because ebg ß-gal would not be able to utilize lactose. Roth et al.50 have suggested a possible sequential selection scheme with the ebgR mutations occurring first (as they are more likely), giving the cell some survival advantage. This is followed by the less frequent mutations in ebgA. However, it is difficult to understand why the individual mutations in either ebgR or ebgA would be positively selected for individually.
Roth et al.50 has suggested that the frequency of either the 92 or 977 mutation in ebgA is 10-9 (thus it would follow that the combined mutation frequency for both of these mutations in ebgA is 10-18). The suggested frequency of a mutation occurring in ebgR is 10-6.50 (These frequencies are based on the spontaneous mutation rate in growing cells not under starvation conditions which may or may not be applicable to nondividing or slowly growing cells under starvation conditions.) Thus, the combined mutation frequency of all three mutations occurring in a single strain would be 10-24. This makes it very improbable that all three mutations would occur by chance in a single bacterial cell in a natural setting. While it is possible to obtain strains with all three mutations, this has only been observed in a laboratory setting using an “intelligent” selection scheme.
More contrived schemes for evolving lactose utilization
In a more recent paper, Hall48 presents new work that he has initiated to predict evolutionary potential by “reevolving” lactose utilization in an E. coli strain that is lacZ- and ebgA-. Previous attempts of this approach have failed.51 This latest attempt was made using a lacZ- and ebgA- stain that also contained activating, constitutive mutations in the cryptic ß-glucoside operons.48 Based on previous work with lac operons in Klebsiella,52 Hall had predicted that the ß-glucosides would mutate and become ß-galactosidases.53 Even if this new mutation scheme works, the cells were engineered with a specific, plausibly helpful mutation before the selection schemes began. It is not known how representative this would be of what could occur naturally.
Limited capabilities of adaptive mutation
As stated previously, the only mutations ever found by Hall in ebgA were changes affecting residues 92 and 977.20 Rather than showing the supposed awesome capabilities of mutation and natural selection as mechanisms for evolution this would seem to indicate that mutation and natural selection are very limited in what they can accomplish. The 92 and 977 changes are within the active site of ebgA, and ebg ß-gal has known homology to other ß-galactosidases.20 It is difficult to fathom the number of mutations that would need to occur in an unrelated enzyme to accomplish the same goal (utilization of lactose). Riley and Anilionis state:
The gain of metabolic function by these mechanisms [adaptive mutation] relies on the presence in the genome of genetic information that is nearly adequate to the task at the time the selection condition is imposed.26 Mechanisms of this kind [generating adaptive mutations] would seem best suited to find tuning and the maintenance of a certain degree of responsiveness, but such mechanisms do not seem suited to the task of bringing about the larger kinds of evolutionary changes that must have been required in the acquisition by primitive organisms of major new metabolic capabilities that would require acquisition of entire new pathways and new forms of energy capture.26
Later work by Hall seems to have born this out as he was unable to “evolve” lactose utilization (“despite extensive efforts”) in an E. coli strain that was lacZ- and ebgA-.51 However, in the same article Hall states, “Obviously, given a sufficient number of substitutions, additions, and deletions, the sequence of any gene can evolve into the sequence of any other gene”.51 This statement seems contradictory to his findings, and seems to be based more on a theoretical possibility, but not probable in reality.
In vitro compartmentalization studies performed using ebgA provide further evidence that while many mutations in genes may be possible (theory), they are not probable (reality).11 These studies utilized a cellfree system of aqueous droplets into which a lacZ-, ebgA/C mutant library had been incorporated.11 The activity of the resultant mutant ebg ß-gals was assessed using a fluorescent substrate.11 Many different mutations in ebgA (in conjunction with mutations in ebgC) resulted in ebg ß-gals that had activity towards the substrate.11 These mutations were typically a combination of multiple base substitutions in ebgA and multiple base substitutions and/or nonsense mutations in ebgC.11 Interestingly, only one Class I mutant was found and no Class II mutants were found.11 While this system is extremely artificial and the activity of the ebgA/C mutants towards lactose is not known, it would seem to indicate that although many mutations in ebgA/C result in active ß-gals that natural selection cannot effectively “see” them. It is plausible that some of these same mutations in ebgA/C occurred in the experimental conditions used by Hall. However, these mutants were not selected by natural selection.
Table 4. Kinetic properties of ebgA and lac ß-galactosidases.Summary of the activities of the wild-type ebgA, ebgA mutants, and lacZ ß-galactosidases with various substrates.
| Enzyme | Lactose | Lactulose | Galactosyl-arabinose | Lactobionate |
|---|---|---|---|---|
| Wild-type ebgA | 1.46 | 0.62 | 0.12 | no detectable activity |
| Class I | 8.2 | 0.16 | 0.43 | no detectable activity |
| Class II | 5.4 | 4.3 | 0.82 | no detectable activity |
| Class IV | 3.4 | 0.99 | 1.7 | 0.24 |
| Class V | 1.4 | 0.49 | 0.80 | 3.0 |
| lacZ | 150 | 19.6 | 37.5 | not determined |
| *Data from Hall.20 | ||||
| *Kcat: moles of substrate hydrolyzed per second per mole of enzyme at 37 °C. | ||||
It is possible that many of the mutations observed by Mastrobattista11 in ebgA/C fall into the “no-selection zone” as hypothesized by Kimura.54 This zone includes mutations which are nearly neutral, and although altering the genotype individually these mutations have no detectable effect on the phenotype. Since selection occurs at the phenotypic level, these mutations are essentially not “seen”.55 While a theoretical plausibility, these mutations are highly improbable.
In vivo, many more constraints exist that may limit the mutations in ebgA to the 92 and 977 residues. For example, the interaction of ebgA/C with other substrates, proteins, or regulatory molecules may be affected by many of the mutations discovered in vitro. The ß subunit encoded by ebgC may play an important role in these interactions since no mutations in this gene are found in strains that survive and grow in vivo but many mutations including nonsense mutations are found in ebgC in vitro.
Hall suggests that possibly the genetic code actually imposes constraints on the changes in the resulting protein (besides the change in the “shape” of the protein).51 Other amino acid replacements at 92 and 977 may allow ebg ß-gal to function effectively; however, these changes may require two mutations versus one.51 The in vitro studies, while confined to the same genetic code, do not face the selective pressures of the in vivo situation. Thus, multiple mutations are more readily observed. Hall suggests,
The present study shows that, independent of that outcome [a proposed amino acid replacement other than the one found], the existing sequence of the wild-type ebgA gene, together with the genetic code, impose an evolutionary constraint against that particular solution under these experimental conditions.51
This further confirms the concept that while many mutations are possible, not all mutations are probable.
If the ebg operon is a backup system for more than just the lac operon, fewer changes may be preferable because nutrient conditions may change. The ebg operon may then need to be altered to allow for the catabolism of a substrate other than lactose. This allows the bacteria to maintain a sort of “rigid flexibility”. Transient mutator populations (such as those proposed by the hypermutable state model for adaptive mutation—see appendix) allow bacterial populations to change rapidly in a constantly fluctuating environment where adaptation is necessary for survival and growth. For bacteria this would appear to be an efficient use of energy and resources. However, as stated previously, adaptive mutations do not create the existence of new functional systems (as required by evolution) but rather allow for the remarkable utilization of current genetic material (as required by adaptation).
Adaptive mutation results in limited changes in protein function
The amino acid replacements at 92 and 977 did not result in ebg ß-gal performing a completely new function. Wild-type ebg ß-gal does have the ability to catabolize lactose, albeit inefficiently (table 4).20 Thus, the mutations in ebgA served to enhance its ability to catabolize lactose compared to wild-type ebgA (table 4).20 Enhancing a previously existing function fits well within the paradigm that adaptive mutations help an organism survive stressful environments. It is not evidence that adaptive mutations play a role in evolution, which requires the gain of completely new functions.
It is a well known fact that mutations typically alter the previously existing function of a protein as has been shown in the case of many antibiotic resistance mutations.2 This appears to be applicable to Hall’s work as well. For example, Class IV mutants, although able to metabolize a wider range of sugars, actually have less activity towards lactose (the only natural sugar used in these experiments) than Class I and II (table 4).20 Additionally, Hall discovered that Class V mutants, which are Class IV mutants that can also catabolize lactobionate, do so at the cost of not being able to catabolize other sugars as well as the Class IV mutants (table 4).20 This does not even take into account the possible alteration of the natural function of ebg ß-gal. Adaptive mutations cannot help an organism move in a vertical direction as required by evolution if the mutations consistently counteract themselves. They can only help in the horizontal direction as is necessary for adaptation to a constantly changing environment.
Adaptive mutation has no foresight
As has been stated previously, the three mutations required for the Ebg system to be independent of the need for IPTG (an artificial inducer); one in ebgR and two in ebgA. No selection scheme could produce all three mutations in a single step. In fact, any one of the mutations would likely be selected against. The bacteria would not keep a single, useless, mutation with the anticipation that future mutations will allow this initial mutation to become beneficial. These mechanisms have no such foresight. This is also true of evolution—it could not keep a single mutation that serves no useful function with the hope that further mutations will allow the initial mutation to provide some benefit in a given environment. Evolution also has no foresight.
Creation Model for Understanding and Utilizing Adaptive Mutation
Classification of bacterial kinds
Adaptive mutations in bacteria are a testament to the grace of God in supplying for His creation. After the Fall and Flood, the conditions of the world changed dramatically. Many organisms did not survive. However, bacteria designed with the ability to adapt to ever changing environments most likely coped well with these changes. One of these mechanisms, adaptive mutation and in particular the hypermutable state model (see appendix), has been described in this work. Other models, such as the amplification model and amplification-mutagenesis model, have also been suggested to lead to adaptive mutation.56, 57 In addition, depending on the starvation condition and the original genotype of the organism, the mechanism employed to adapt may differ. For example, during amino acid starvation E. coli derepresses the biosynthesis operon for the specific limited amino acid. Wright, Longacre, and Reimers58 found that in leuB- E. coli the leu operon is de-repressed and E. coli can revert to leuB+ by a starvationinduced transcription model. This leads to increased transcription of only the leu operon and hypermutation specifically in the leu operon.58 This is different from the hypermutable state model proposed by Hall for the Ebg system where genome wide hypermutation is observed during lactose starvation (appendix).42, 6 Determining the adaptive mechanisms used by the spectra of bacterial species may assist in the process of classification of bacterial kinds. To date, no attempt has been made to classify bacteria into original created kinds.
Pathogenesis
It is likely that adaptive mutation was not needed until after the Fall, when bacteria began dealing with an imperfect environment. Mutations in bacteria that lead to pathogenicity would not have occurred until after the Fall. One possibility is that pathogenicity is a detrimental (to other organisms) side effect of adaptive mutation. Pathogenicity may not be the result of completely random mutations. Joseph Francis59 suggests,
From a creation view, it appears then, that the origin of microbial based disease has at least two primary causes (1) post-Fall genetic alteration of the original good microbe and/or (2) post-Fall displacement or movement of the microbe from the site where it performed its beneficial function.
Adaptive mutations (“alteration[s]”) while performing a beneficial function in a given environment may have pathogenic effects should the microbe move or be moved to a different location (“displacement”). This phenomenon of displacement is observed when bacteria on the skin (which have no pathogenic effect) enter the blood stream through a break in the skin (such as during surgery) and exert a pathogenic effect. As more pathogenic bacterial genomes are sequenced, it will be important to understand the function of the proposed pathogenic genes in the metabolic activities of the bacteria.
Limits of genetic changes
Neo-Darwinism proposes that mutation and natural selection make completely random changes in organisms that over time lead to a microbe becoming a microbiologist. Adaptive mutation throws a wrench into the evolution machine. Wright, Longacre, and Reimers state,
The current paradigm of neo-Darwinism as formulated by Weisman rejects any influence of the environment on the direction of variation. However, prolonged nutritional stress results in a general increase in mutations rates; the introduction of environmental effects on specific mutations rates is a reasonable extension of what is known . . . .58
The body of evidence seems to indicate that adaptive mutation is a real phenomenon.57 For creationists, adaptive mutation fits well within the paradigm of God’s design and providence for His creation. God foreknew that the Fall and Flood would occur and that bacteria would need the ability to adapt to altered environments. Bacteria have been designed with great genetic diversity that sometimes is only accessible through genetic alterations such as adaptive mutations. Thus, the ebg operon may be part of a complex backup system for the lac operon in E. coli.
Bacteria are great economists and change according to supply and demand. Random changes that do not help them deal with their environment will likely be lost quickly. Adaptive mutations provide an even greater genetic flexibility and response. A logical assumption, therefore, is that bacteria were created with an inherent ability for adapting to rapid and dramatic environmental changes. As such, adaptive mutation may be the reigning mechanism for change in bacteria, especially considering they typically reside in nutrient poor environments. Adaptive mutation alters current functional systems in bacteria; it does not serve to add new functional systems necessary for evolution. Thus it is an error to assume such mutations can produce large genetic changes in the bacterium (such as those necessary for “common descent”). Certainly, adaptive mutations fit well with a creation model that allows for change within a created kind (or baramin), but they do not appropriately fit with change between created kinds.
Conclusion
Rather than being an example of “evolution in action,” adaptive mutation is an awesome witness to God’s design of bacteria. Hall’s discovery of adaptive mutations in the Ebg system do not pass Miller’s8 “true acid test” for the evolution of a new biochemical pathway. Evolution requires random mutations, natural selection, and time to bring about new functional systems. Mutations in the ebg operon do not serve to add a new functional system to the bacteria. Rather they enhance a previously existing function of ebg ß-gal, that of catabolizing lactose. Adaptive mutation leads to the alteration of current genetic material to allow the bacteria to adapt to adverse environmental conditions. The terms evolution and adaptation should not be equivocated. Adaptive mutations are not evidence for evolution (defined as descent with modification) as the bacteria remain bacteria with the gain and loss of specific functions. Although the mutations in the ebg operon were obtained through contrived methods that may not occur naturally, Hall’s work has helped to show what bacteria are potentially capable of and that adaptive mutation is a real phenomenon.
Acknowledgments
We would also like to thank Barry Hall for communications with us that provided an inside look at his research.
Appendix: Possible Mechanisms for Generating Adaptive Mutations
The mechanisms for generating adaptive mutations are still being debated. It appears there is more than one mechanism to accomplish the same goal, and the mechanism(s) employed depends on the species and the starvation conditions. Tenallion, Taddei, Radman, and Matic suggest a type of second-order selection applies here:
. . . while selecting for adaptive mutations [first-order selection], evolution [sic] indirectly selects for a system that creates these adaptive mutations, thus allowing second-order selection to regulate the mutational process. This results in at least transient enrichment for cells exhibiting increased rates of genetic change.60
Under starvation conditions, a stress response is initiated in bacteria (fig. 3). The first step in the stress response is the production of the sigma factor, ss (encoded by the rpoS gene).61 ss binds to RNA polymerase and affects which promoters are recognized and transcribed and also plays a role in translation efficiency and protein stability.61 ss is also believed to play a role in making double-strand-break repair error prone in multiple ways.62 There are several proposed models to explain pathways leading to adaptive changes following rpoS induction. The pathway choice may be dependent on the bacterial species, starvation conditions, and/or initial mutations in the bacterial population being studied.
Hypermutation model
The hypermutation model (also called hypermutable state model) is the most applicable to Hall’s research on the ebg operon (fig. 3). Hall was the first to suggest the idea of hypermutation in bacterial colonies undergoing starvation.
Consider that, at any instant during periods of prolonged starvation, some fraction of the cells in a colony enter into a hypermutable state in which extensive DNA damage and resulting error-prone repair synthesis occur, while the remaining cells in the colony remain essentially immutable. When cells are in the hypermutable state, mutations might occur at many sites in the same cell. If one of those mutations provided a solution to the current problem of blocked growth . . . then the cell could exit the hypermutable state and be recovered (as a mutant).42
It appears that genome wide hypermutation occurs in a subpopulation of the cells being subjected to starvation/selection. These cells in addition to adaptive mutations which allow them to grow also have mutations in genes not under selection.6 Other studies have shown that most of these cells are in a transient state of hypermutation versus a heritable one.6 The advantage of this would be that once a mutation occurs that allows growth in the given conditions, further mutations and subsequent damage to the genome would not occur due to exiting the hypermutable state. The state of hypermutation can be achieved through multiple mechanisms that are all likely initiated by ss. All of these mechanisms are related to DNA replication, proofreading and repair.63 Although it seems counterintuitive that nondividing or slowly dividing cells under starvation would be synthesizing much DNA, there is evidence that DNA synthesis occurs at 0.5%–5% per genome per day.61
Stress is believed to cause double stranded breaks (DSB) in DNA. DSBs activate the SOS response in bacteria. This induces the expression of Pol IV (encoded by the gene DinB).64 ss also increases the expression of DinB.62 Pol IV is an error-prone polymerase that causes frameshifts and substitutions during DNA replication.64 The large number of mutations generated by Pol IV DNA synthesis effectively saturates the mismatch repair system leading to a certain number of mutations escaping DNA repair and thus, genome wide hypermutation.62 Once the adaptive mutation(s) has occurred growth begins and the stress response ceases.
In the case of Hall’s work on the ebg operon, mutations in both ebgR and ebgA would need to occur before exiting the hypermutable state. Mutations in ebgR are fairly common (around 10-6—as many mutations will disable the repressor) whereas mutations in ebgA are much rarer (around 10-18—two changes at positions 92 and 977 are necessary to allow utilization of lactose).65 (These frequencies are based off the spontaneous mutation rate in growing cells not under starvation conditions.) Some have questioned if the mutation rate necessary for a workable hypermutation model is unrealistic.65 Mutation rates for growing cells may not be applicable to starved cells. Mechanisms for generating mutations in growing cells are different than those in starved cells evidenced by the ability of certain mutant strains to revert when growing but not under prolonged selection.66
The specificity of the mutations seen in ebgA appears to be the result of selective capture vs. selective generation. Although selective generation has not been completely ruled out, it seems unlikely that a mechanism exists which specifically targets genes under selection.66 Rather than selection generating mutations, selection is capturing mutations that give cells a growth advantage under the selection conditions.66 While mutations in genes not under selection occur, they are likely to be “hitchhikers” resulting from genome wide hypermutation.
Footnotes
- Hoekstra, H. E., R. J. Hirschmann, R. A. Bundey, P. A. Insel, and J. P. Crossland, 2006. A single amino acid mutation contributes to adaptive beach mouse color pattern. Science 313:101–104.
- Anderson, K. L., 2005. Is bacterial resistance to antibiotics an appropriate example of evolutionary change? Creation Research Science Quarterly 41:318–326.
- Bjedov, I., O. Tenaillon, G. Bénédicte, V. Souza, E. Denamur, M. Radman, et al., 2003. Stress-induced mutagenesis in bacteria. Science 300:1404–1409.
- Hall, B. G., 1991. Adaptive evolution that requires multiple spontaneous mutations: Mutations involving base substitutions. Proceedings of the National Academy of Sciences 88:5882–5886.
- Hall, B. G., and D. L. Hartl, 1974. Regulation of newly evolved enzymes. I. Selection of a novel lactase regulated by lactose in Escherichia coli. Genetics 76:391–400.
- Torkelson, J., R. S. Harris, M-J. Lombardo, J. Nagendran, C. Thulin, and S. M. Rosenberg, 1997. Genome-wide hypermutation in a subpopulation of stationary-phase cells underlies recombination-dependent adaptive mutation. The EMBO Journal 16: 3303–3311.
- Behe, M., 1998. Darwin’s black box: The biochemical challenge to evolution. New York, New York: The Free Press.
- Miller, K. R., 1999. Finding Darwin’s God. New York, New York: HarperCollins Publishers, p. 145.
- Behe, M., 2000. A true acid test: Response to Ken Miller. Retrieved October, 18, 2006, from Access Research Network web site: http://www.arn.org/docs/behe/mb_trueacidtest.htm.
- Miller, K. R. (n.d.). “A true acid test”. Retrieved October 24, 2006 from http://www.millerandlevine.com/km/evol/DI/AcidTest.html.
- Mastrobattista, E., V. Taly, E. Chanudet, P. Treacy, B. T. Kelly, and A. D. Griffiths, 2005. High-throughput screening of enzyme libraries: Evolution of a ß-galactosidase by fluorescence-activated sorting of double emulsions. Chemistry & Biology 12:1291–1300.
- Anderson, K. L., 2003. The complex world of gastrointestinal bacteria. Canadian Journal of Animal Science 83:409—427.
- Frair, W., 2000. Baraminology—classification of created organisms. Creation Reseach Society 37:82–91.
- Hall, B. G., 1998b. Adaptive mutagenesis at ebgR is regulated by PhoPQ. Journal of Bacteriology 180:2862.
- Foster, P. L., 1999. Mechanisms of stationary phase mutation: A decade of adaptive mutation. Annual Review of Genetics 33: 57–88.
- Cairns, J., and P. L. Foster, 1991. Adaptive reversion of a frameshift mutation in Escherichia coli. Genetics 128:695–701.
- Hall, B. G., 1978a. Regulation of newly evolved enzymes. IV. Directed evolution of the ebg repressor. Genetics 90:673–681.
- Hall, B. G., 1978b. Experimental evolution of a new enzymatic function. II Evolution of multiple functions for ebg enzyme in E. coli. Genetics 89:453–465.
- Hall, B. G., and D. L. Hartl, 1975. Regulation of newly evolved enzymes II. The ebg repressor. Genetics 81:427–435.
- Hall, B. G., 1999a. Experimental evolution of Ebg enzyme provides clues about the evolution of catalysis and to evolutionary potential. FEMS Microbiology Letters 174:1–8.
- Hall, B. G., P. W. Betts, and J. C. Wootton, 1989. DNA sequence analysis of artificially evolved ebg enzyme and ebg repressor genes. Genetics 123:635–648.
- Hall, B. G., and N. D. Clarke, 1977. Regulation of newly evolved enzymes. III Evolution of the ebg repressor during selection for enhanced lactase activity. Genetics 85:193–201.
- Stokes, H. W., P. W. Betts, and B. G. Hall, 1985. Sequence of the ebgA gene of Escherichia coli: Comparison with the lacZ gene. Molecular Biology and Evolution 2:469–477.
- Stokes, H. W., and B. G. Hall, 1985. Sequence of the ebgR gene of Escherichia coli: Evidence that the EBG and LAC operons are descended from a common ancestor. Molecular Biology and Evolution 2:478–483.
- Kellis, M., B. W. Birren, and E. S. Lander, 2004. Proof and evolutionary analysis of ancient genome duplication in the yeast Saccharomyces cerevisiae. Nature 428:617–624.
- Riley, M., and A. Anilionis, 1978. Evolution of the bacterial genome. Annual Reviews of Microbiology 32519–560.
- Hall, B. G., 1999b. Spectra of spontaneous growth-dependent and adaptive mutations at ebgR. Journal of Bacteriology 181:1149–1155.
- Bergman, J., 2006. Does gene duplication provide the engine for evolution? Journal of Creation 20:1, 99–104.
- Hall, B. G., 1981. Changes in the substrate specificities of an enzyme during directed evolution of new function. Biochemistry 20:4042–4049.
- Hall, B. G., and T. Zurel, 1980. Evolution of a new enzymatic function by recombination within a gene. Proceedings of the National Academy of Sciences 77:3529–3533.
- Rolseth, S. J., V. A. Fried, and B. G. Hall, 1980. A mutant ebg enzyme that converts lactose into an inducer of the lac operon. Journal of Bacteriology 142:1036–1039.
- Stoebel, D. M., 2004. Lack of evidence for horizontal transfer of the lac operon into Escherichia coli. Molecular Biology and Evolution 22:683–690.
- Herring, C. D., A. Raghunathan, C. Honisch, T. Patel, M. K. Applebee, A. R. Joyce, A. R., et al., 2006. Comparative genome sequencing of Escherichia coli allows observation of bacterial evolution on a laboratory timescale. Nature Genetics 38: 1406–1412.
- Liu, Y., P. M. Harrison, V. Kunin, and M. Gerstein, 2004. Comprehensive analysis of pseudogenes in prokaryotes: Widespread gene decay and failure of putative horizontally transferred genes. Genome Biology 5, Article R64. Retrieved April 19, 2007, from https://genomebiology.biomedcentral.com/articles/10.1186/gb-2004-5-9-r64.
- Balakirev, E. S., and F. J. Ayala, 2003. Pseudogenes: Are they “junk” or functional DNA? Annual Review of Genetics 37: 123–151.
- Zheng, D., A. Frankish, R. Baertsch, P. Kapranov, A. Reymond, S. W. Choo, Y. Lu, F. Denoeud, S. E. Antonarakis, M. Snyder, Y. Ruan, C. L. Wei, T. R. Gingeras, R. Guigó, J. Harrow, and M. B. Gerstein, 2007. Pseudogenes in the ENCODE regions: Consensus annotation, analysis of transcription, and evolution. Genome Research 17:839–851.
- Pál, C., B. Papp, M. J. Lercher, P. Csermely, S. G. Oliver, and L. D. Hurst, 2006. Chance and necessity in the evolution of minimal metabolic networks. Nature 440:667–670.
- Kafri, R., M. Levy, and Y. Pilpel, 2006. The regulatory utilization of genetic redundancy through responsive backup circuits. Proceedings of the National Academy of Sciences 103:11653–11658.
- García-Rodriguez, L. J., J. A. Trilla, C. Castro, M. H. Valdivieso, A. Durán, and C. Roncero, 2000. Characterization of the chitin biosynthesis process as a compensatory mechanism in the fks1 mutant of Saccharomyces cerevisiae. FEBS Letters 478:84–88.
- Groisman, E. A., F. Hefron, and F. Solomon, 1992. Molecular genetic analysis of the Escherichia coli phoP locus. Journal of Bacteriology 174:486–491.
- Cairns, J., J. Overbaugh, and S. Miller, 1988. The origin of mutants. Nature 335:142–145.
- Hall, B. G., 1990. Spontaneous point mutations that occur more often when advantageous than when neutral. Genetics 126:5–16.
- Mayr, E., 1982. The growth of biological thought: Diversity, evolution and inheritance. Cambridge, Massachusetts: Harvard University Press.
- Stahl, F. W., 1988. A unicorn in the garden. Nature 335:112–113.
- Lenski, R. E., M. Slatkin, and F. J. Ayala, 1989. Mutation and selection in bacterial populations: Alternatives to the hypothesis of directed mutation. Proceedings of the National Academy of Sciences, USA 86:2775.
- Anderson, K. L. and G. Purdom, 2008. A creationist perspective of beneficial mutations in bacteria. In A. A. Snelling, (ed.), Proceedings of the sixth international conference on creationism. Pittsburgh, Pennsylvania: Creation Science Fellowship & Dallas, Texas: Institute for Creation Research (pp. 73–86).
- Hall, B. G., 1988. Adaptive evolution that requires multiple spontaneous mutations. I. Mutations involving an insertion sequence. Genetics 120:896.
- Hall, B. G., 2003. The EBG system of E. coli: Origin and evolution of a novel ß-galactosidase for the metabolism of lactose. Genetics 118:143.
- Lenski, R. E., and M. Travisano, 1994. Dynamics of adaptation and diversification: A 10,000-generation experiment with bacterial populations. Proceedings of the National Academy of Science, USA:91:6808–6814.
- Roth, J. R., E. Kugelberg, A. B. Reams, E. Kofoid, and D. I. Andersson, 2006. Origin of mutations under selection: The adaptive mutation controversy. Annual Review of Microbiology 60:477–501.
- Hall, B. G., 1995. Evolutionary potential of the ebgA gene. Molecular Biology of Evolution 12:514–517.
- Hall, B. G., 1979. Lactose metabolism involving phosph-ß-galactosidase in Klebsiella. Journal of Bacteriology 138:691–698.
- Hall, B. G., 2001. Predicting evolutionary potential. I. Predicting the evolution of a lactose-PTS system in Escherichia coli. Molecular Biology and Evolution 18:1389–1400.
- Kimura, M., 1979. Model of effective neutral mutations in which selective constraint is incorporated. Proceedings of the National Academy of Sciences, USA 76:3440–3444.
- Sanford, J. C., 2005. Genetic entropy and the mystery of the genome. Lima, New York: Ivan Press.
- Hendrickson, H., E. S. Slechta, U. Bergthorsson, D. I. Andersson, and J. R. Roth, 2002. Amplification-mutagenesis: Evidence that “directed” adaptive mutation and general hypermutability result from growth with a selected gene amplification. Proceedings of the National Academy of Sciences 99:2164–2169.
- Stumpf, J. D., A. R. Poteete, and P. L. Foster, 2007. Amplification of lac cannot account for adaptive mutation to Lac+ in Escherichia coli. Journal of Bacteriology 189:2291–2299.
- Wright, B. E., A. Longacre, and J. M. Reimers, 1999. Hypermutation in derepressed operons of Escherichia coli K12. Proceedings of the National Academy of Sciences 96:5089–5094.
- Francis, J. W., 2008. Creation, microbiology and the origin of disease. In Purdom, G., and J. W. Francis (eds.), Proceedings of the Microbe Forum, June 2007. Answers Research Journal 1:1–6.
- Tenaillon, O., F. Taddei, M. Radman, and I. Matic, 2001. Second-order selection in bacterial evolution: Selection acting on mutation and recombination rates in the course of adaptation. Research in Microbiology 152:11.
- Zambrano, M. M., and R. Kolter, 1996. GASPing for life in stationary phase. Cell 86:181–184.
- Rosenberg, S. M., and P. J. Hastings, 2004. Adpative point mutation and adaptive amplification pathways in the Escherichia coli lac system: Stress responses producing genetic change. Journal of Bacteriology 186:4838–4843.
- Bridges, B. A., 2001. Hypermutation in bacteria and other cellular systems. Philosophical Transactions of The Royal Society B 356:29–39.
- McKenzie, G. J., and S. M. Rosenberg, 2001. Adaptive mutations, mutator DNA polymerases and genetic change strategies of pathogens. Current Opinion in Microbiology 4:586–594.
- Roth, J. R., E. Kofoid, F. P. Roth, O. G. Berg, J. Seger, and D. I. Andersson, 2003. Regulating general mutation rates: Examination of the hypermutable state model for Cairnsian adaptive mutation. Genetics 163:1483–1496.
- Hall, B. G., 1998a. Adaptive mutagenesis: A process that generates almost exclusively beneficial mutations. Genetica 102/103:109–125.

Answers in Genesis is an apologetics ministry, dedicated to helping Christians defend their faith and proclaim the good news of Jesus Christ.
- Customer Service 800.778.3390
- Available Monday–Friday | 9 AM–5 PM ET
- © 2026 Answers in Genesis

